Towards Automatic Face-to-Face Translation
ACM Multimedia 2019
[Code]
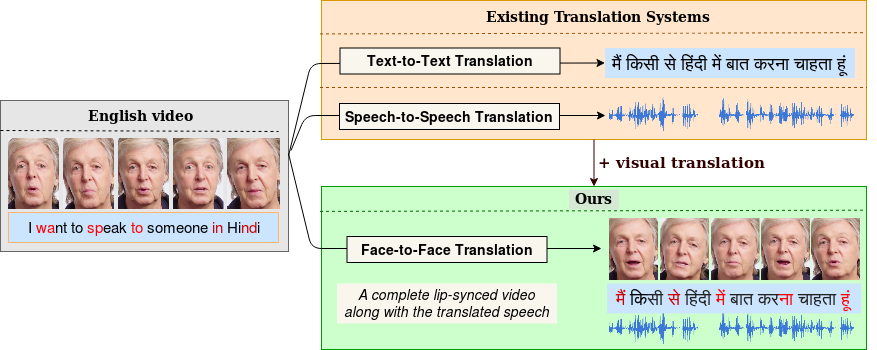
Given a speaker speaking in a language L$_A$ (Hindi in this case), our fully-automated system generates a video of the speaker speaking in L$_B$ (English). Here, we illustrate a potential real-world application of such a system where two people can engage in a natural conversation in their own respective languages.
Abstract
In light of the recent breakthroughs in automatic machine translation systems, we propose a novel approach of what we term as "Face-to-Face Translation". As today's digital communication becomes increasingly visual, we argue that there is the need for systems that can automatically translate a video of a person speaking in language A into a target language B with realistic lip synchronization. In this work, we create an automatic pipeline for this problem and demonstrate its impact in multiple real-world applications. First, we build a working speech-to-speech translation system by bringing together multiple existing modules from speech and language. We then move towards "Face-to-Face Translation" by incorporating a novel visual module, LipGAN for generating realistic talking faces from the translated audio. Quantitative evaluation of LipGAN on the standard LRW test set, shows that it significantly outperforms existing approaches across all standard metrics. We also subject our Face-to-Face Translation pipeline, to multiple human evaluations and show that it can significantly improve the overall user experience for consuming and interacting with multimodal content across languages.
Paper
-

Towards Automatic Face-to-Face Translation
Prajwal Renukanand*, Rudrabha Mukhopadhyay*, Jerin Philip, Abhishek Jha, Vinay Namboodiri and C.V. Jawahar
Towards Automatic Face-to-Face Translation, ACM Multimedia, 2019.
[PDF] | [BibTeX]@inproceedings{KR:2019:TAF:3343031.3351066,
author = {K R, Prajwal and Mukhopadhyay, Rudrabha and Philip, Jerin and Jha, Abhishek and Namboodiri, Vinay and Jawahar, C V},
title = {Towards Automatic Face-to-Face Translation},
booktitle = {Proceedings of the 27th ACM International Conference on Multimedia},
series = {MM '19},
year = {2019},
isbn = {978-1-4503-6889-6},
location = {Nice, France},
= {1428--1436},
numpages = {9},
url = {http://doi.acm.org/10.1145/3343031.3351066},
doi = {10.1145/3343031.3351066},
acmid = {3351066},
publisher = {ACM},
address = {New York, NY, USA},
keywords = {cross-language talking face generation, lip synthesis, neural machine translation, speech to speech translation, translation systems, voice transfer},
} }
Demo
<!<!
Click here to redirect to the video
Speech-to-Speech Translation
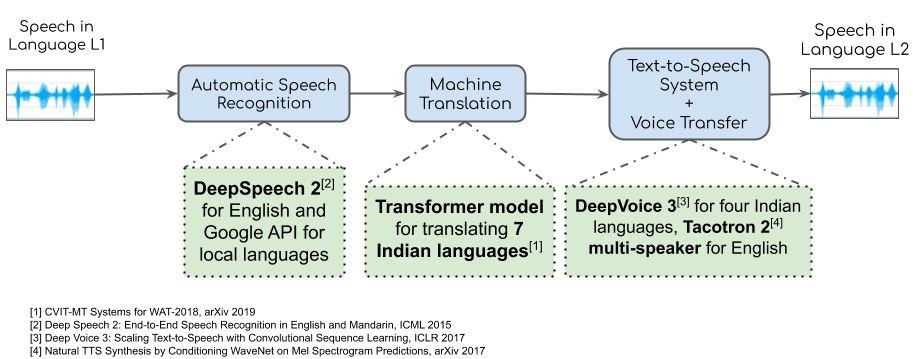
Pipeline for Speech-to-Speech Translation
Our system can be widely divided into two sub-systems, (a) Speech-to-Speech Translation and (b) Lip Synthesis. We do speech-to-speech translation by combining ASR, NMT and TTS. We first use a publicly available ASR to get the text transcript. For English we use DeepSpeech for transcribing English text from audio. We use a suitable publicly available ASR for other languages like Hindi and French. We train our own NMT system for different Indian languages using Facebook AI Research's publicly available codebase. We finally train a TTS for each language of our choice. The TTS is used to generate speech in the target language.
Synthesizing Talking Faces from Speech
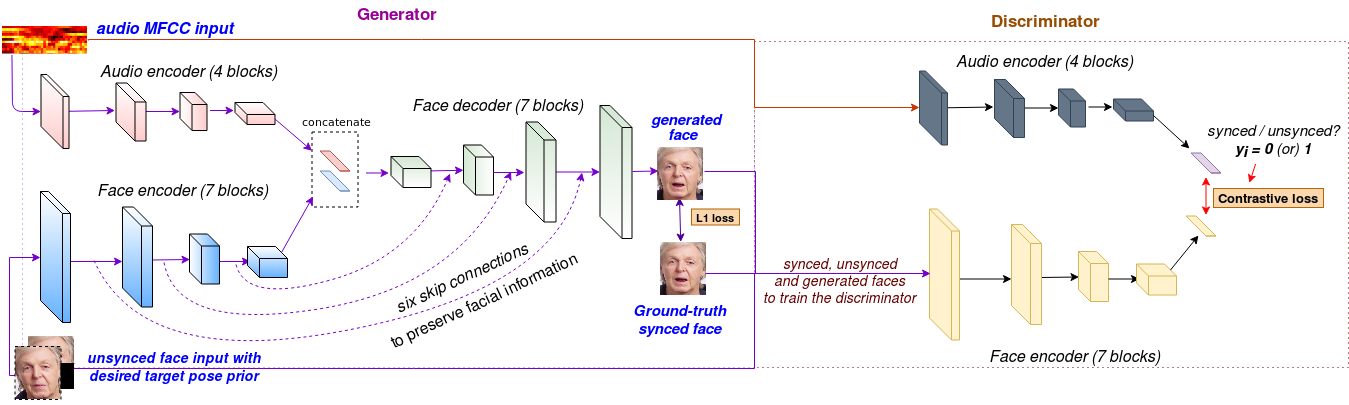
Pipeline for generating talking faces of any identity given a speech segment
We create a novel model called LipGAN which can generate talking faces of any person given a speech segment. The model comprises of two encoders, (a) Face encoder and (b) Speech encoder. The face encoder is used to encode information about the identity of the talking face. The speech encoder takes a very small speech segment (350 ms of audio at a time) and is used to encode the audio information. The outputs from both of these encoders are then fed to a decoder which generates a face image of the given identity which matches the lip shape corresponding to the given audio segment. For more information about our model please go through the paper.