Transfer Learning for Scene Text Recognition in Indian Languages
Sanjana Gunna, Rohit Saluja, and C.V. Jawahar
[Video] [Paper] [ Code]
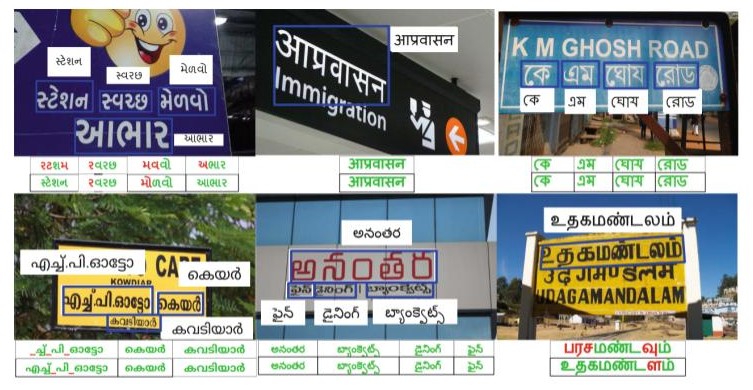
Clockwise from top-left; “Top: Annotated Scene-text images, Bottom: Baselines’ predictions (row-1) and Transfer Learning models’ predictions (row-2)”, from Gujarati, Hindi, Bangla, Tamil, Telugu and Malayalam. Green, red, and “ ” represent correct predictions, errors, and missing characters, respectively. (Color figure online)
Abstract
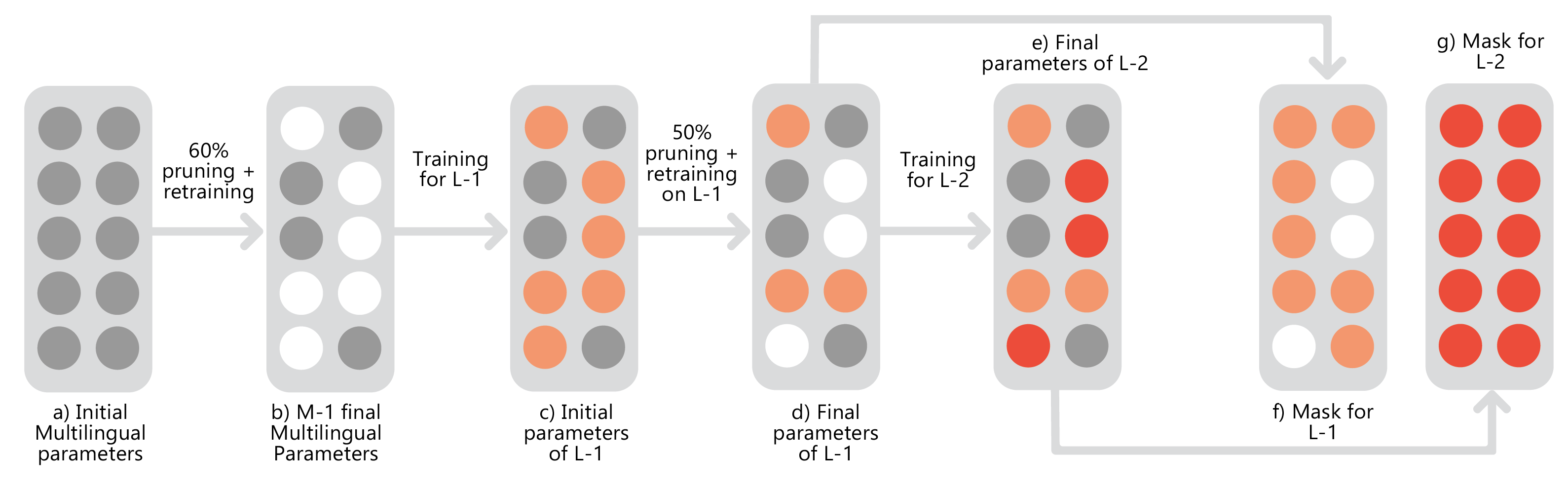
Scene text recognition in low-resource Indian languages is challenging because of complexities like multiple scripts, fonts, text size, and orientations. In this work, we investigate the power of transfer learning for all the layers of deep scene text recognition networks from English to two common Indian languages. We show that the transfer of English models to simple synthetic datasets of Indian languages is not practical. Instead, we propose to apply transfer learning techniques among Indian languages and study the transfer learning among six Indian languages with varying complexities in fonts and word length statistics. We achieve gains in Word Recognition Rates (WRRs) on IIIT-ILST Hindi, Telugu, and Malayalam datasets in comparison to previous works. On the MLT-19 Hindi and Bangla datasets and our Gujarati and Tamil datasets, we observe similar increase in WRR respectively, over our baseline models
We additionally release a dataset of around 440 scene images containing 500 Gujarati and 2535 Tamil words.
Paper
-
Transfer Learning for Scene Text Recognition in Indian Languages
Sanjana Gunna, Rohit Saluja, C.V. Jawahar
Transfer Learning for Scene Text Recognition in Indian Languages, ICDAR 2021 WORKSHOP ON CAMERA-BASED DOCUMENT ANALYSIS AND RECOGNITION (CBDAR 2021, 9TH EDITION), 2021.
[PDF] | [BibTeX]@inproceedings{Gunna2021TransferLF,title={Transfer Learning for Scene Text Recognition in Indian Languages},author={Sanjana Gunna and Rohit Saluja and C. V. Jawahar},booktitle={ICDAR Workshops},year={2021}}
Dataset
The dataset contains nearly 440 real scene images, 500 cropped scene images for Gujarati script and 2535 for Tamil script and are annotated for scene text bounding boxes and transcriptions.
Terms and Conditions: All images provided as part of the dataset have been collected from freely accessible internet sites. As such, they are copyrighted by their respective authors. The images are hosted in this site, physically located in India. Specifically, the images are provided with the sole purpose to be used for research in developing new models for scene text recognition You are not allowed to redistribute any images in this dataset. By downloading any of the files below you explicitly state that your final purpose is to perform research in the conditions mentioned above and you agree to comply with these terms and conditions.
Download
Contact
- Sanjana Gunna -
This email address is being protected from spambots. You need JavaScript enabled to view it. - Rohit Saluja -
This email address is being protected from spambots. You need JavaScript enabled to view it.